Home

The Mohlke Lab performs human genetics research in the Department of Genetics at the University of North Carolina at Chapel Hill. We are affiliated with the Computational Medicine Program, the McAllister Heart Institute, and the Lineberger Comprehensive Cancer Center. PhD students in the Mohlke Lab are part of the Genetics and Molecular Biology (GMB) Curriculum or the Bioinformatics and Computational Biology (BCB) PhD Curriculum, part of the Biological and Biomedical Sciences Program (BBSP).
Research
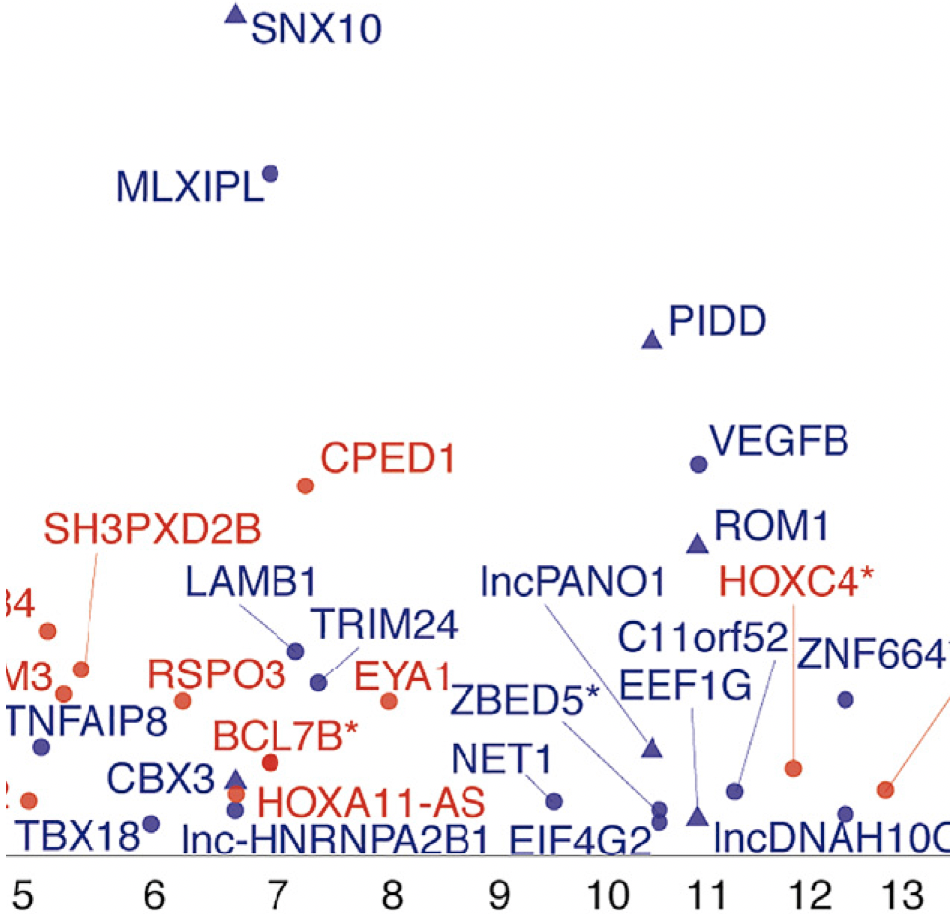
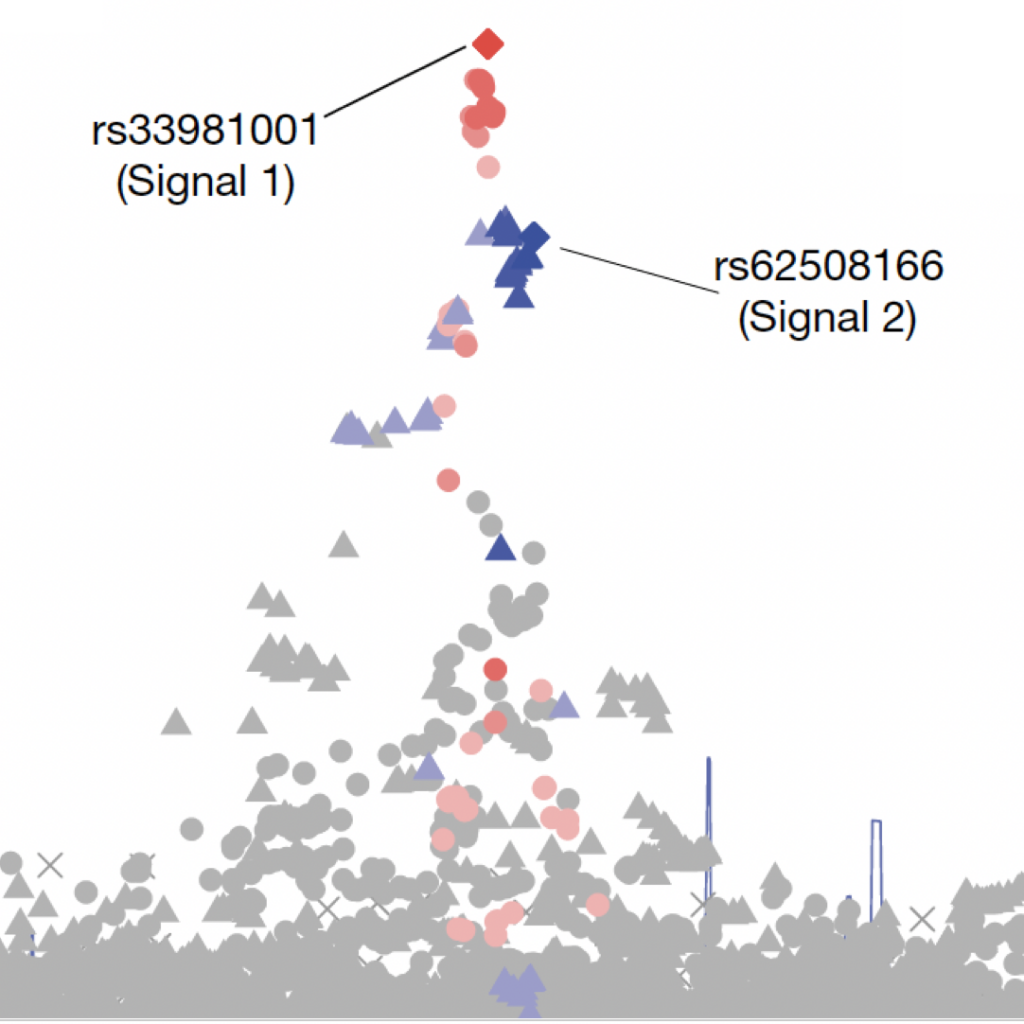


Complex Trait Genetics
We identify genetic loci associated with type 2 diabetes, obesity, dyslipidemia and related quantitative traits.
Regulatory Genomics
We study genetic and environmental effects on epigenome and transcriptome variation in disease-relevant tissues
Disease Mechanisms
We determine how genetic variants affect genes and how genes function to influence disease processes
Recent News
January 2024 Liver regulatory mechanisms paper in HGG Advances
Gautam Pandey, with contributions from Rani, Kevin, Shelley, Jayna, Jessica, and Alaine, experimentally validated regulatory activity at chromatin accessibility QTL and GWAS loci using CRISPRi and transcriptional assays.
November 2023 Adipose cell type composition paper published in Diabetes
Sarah Brotman showed that associations between adipose gene expression levels and cardiometabolic traits are influenced by cell type composition and body mass index.
September 2023 First genome-wide association study for hidradenitis suppurativa
Alaine Broadaway co-first authored, with Quan Sun, the first GWAS paper for this common skin disorder, a collaboration with Christopher Sayed, Yun Li and others.
June 2023 New loci associated with insulin-stimulated glucose uptake
Alaine Broadaway, Shelley Moxley, Rani Vadlamudi and Emma Wilson contributed to discovery of loci for postprandial insulin resistance and showed a potential regulatory molecular mechanism at SLC2A4
March 2023 Congratulations Dr. Brotman!
Sarah Brotman successfully defended her PhD thesis in Genetics and Molecular Biology and earned a certificate in Bioinformatics and Computational Biology. Sarah described the effects of adipose tissue gene expression, isoform usage, and cell type composition on cardiometabolic traits.
March 2023 Foundation of the NIH award
With MPI Laura Scott, we will generate single nucleus RNA-seq and metabolomics data in subcutaneous adipose tissue and identify cell type expression and metabolite quantitative trait lloci
February 2023 New genetic loci for proinsulin levels published in AJHG
Alaine Broadaway, Emma Wilson, Victoria Parsons and coauthors identified and characterized loci in the largest genetic study of proinsulin levels. Shelley Moxley and Rani Vadlamudi showed a plausible islet regulatory mechanism at MADD
September 2022 Sarah wins Magnuson award
Congratulations to Sarah Brotman, who received a Terry Magnuson award given to outstanding graduating students in the Curriculum in Genetics and Molecular Biology
August 2022 SARS-CoV-2 receptor ACE2 expression levels and cardiometabolic risk
Sarah Brotman and coauthors showed that adipose expression level of ACE2 is associated with cardiometabolic trait levels and adipose cell type proportions
June 2022 Welcome new PhD students!
Welcome Abdalla Alkhawaja, a Bioinformatics and Computational Biology student co-mentored by Terry Furey; Jayna Nicholas, a Genetics and Molecular Biology student co-mentored by Laura Raffield; and Sarah Lester, a Genetics and Molecular Biology student co-mentored by Samir Kelada
May 2022 New R01: Genetics of adipose cell-type expression and cardiometabolic traits
With MPIs Paivi Pajukanta and Laura Scott, we will identify cell-type-limited expression quantitative trait loci, adipose genes relevant to cardiometabolic traits, and genetic variants responsible for cell type effects
April 2022 New R01: Regulatory genomics of ozone air pollution response
With MPI Samir Kelada, we will identify gene-by-environment interactions in response of airway brochial epithelial cells to the air pollutant ozone
February 2022 Congratulations Dr. Perrin!
Hannah Perrin successfully defended her PhD thesis in Genetics and Molecular Biology and will also earn a certificate in Bioinformatics and Computational Biology. Hannah described context-dependent regulatory mechanisms for cardiometabolic traits
January 2022 Welcome new postdoc Jon Rosen
Postdoc Jon Rosen, co-mentored by Mike Love, will lead analyses and methods development in the UNC Impact of Genomic Variation on Function project
January 2022 Adipose spliceQTL paper published in AJHG
Sarah Brotman and Chelsea Raulerson described splice junction QTL in adipose tissue, including sQTL that colocalize with cardiometabolic GWAS signals, including at NR1H3. See the article here
January 2022 Gautam awarded AHA postdoc fellowship
Gautam Pandey was awarded a fellowship from the American Heart Association to identify genetic effects on hepatic lipid metabolism and cardiovascular disease
December 2, 2021 One of Sarah Brotman’s outreach activities
The Shadow a Scientist program highlighted Sarah’s role as an ambassador
November 12, 2021 Chancellor’s Science Scholar research profile
Undergraduate Abi Rohy’s research was highlighted by the Chancellor’s Science Scholar program. See the article here.
October 26, 2021 PLoS Genetics paper: Chromatin accessibility during adipocyte differentiation
Hannah Perrin and Kevin Currin described changes of accessible chromatin and gene expression levels during adipocyte differentiation. They identified GWAS variants in differentially accessible regions and demonstrated context-dependent effects on transcriptional activity.
September 1, 2021 Impact of Genomic Variation on Function
With MPIs Hyejung Won and Michael Love, we will join NHGRI’s new IGVF consortium as a characterization center. We will investigate the regulatory consequences of GWAS variants, examining effects of tissue, sex, and environmental perturbation. Many other collaborators include Yun Li and Bev Koller in the UNC Department of Genetics.
May 31, 2021 Glycemic trait GWAS published in Nature Genetics
With many co-authors from MAGIC and co-first author Cassie Spracklen, we identified and characterized loci for fasting glucose, fasting insulin, 2-hour glucose and hemoglobin A1c through trans-ancestry GWAS meta-analysis.
May 25, 2021 Liver chromatin accessibility QTL paper published in AJHG
Kevin Currin described liver caQTL, linked peaks to genes, and performed colocalization with eQTL and GWAS loci. Collaborators include Michael Erdos and Francis Collins at NHGRI.
mohlke@med.unc.edu
Phone
919-966-2913
Office
5096 Genetic Medicine Building
Chapel Hill, NC 27599-7264