Research
Identify Genetic Loci Associated with Complex Diseases and Traits
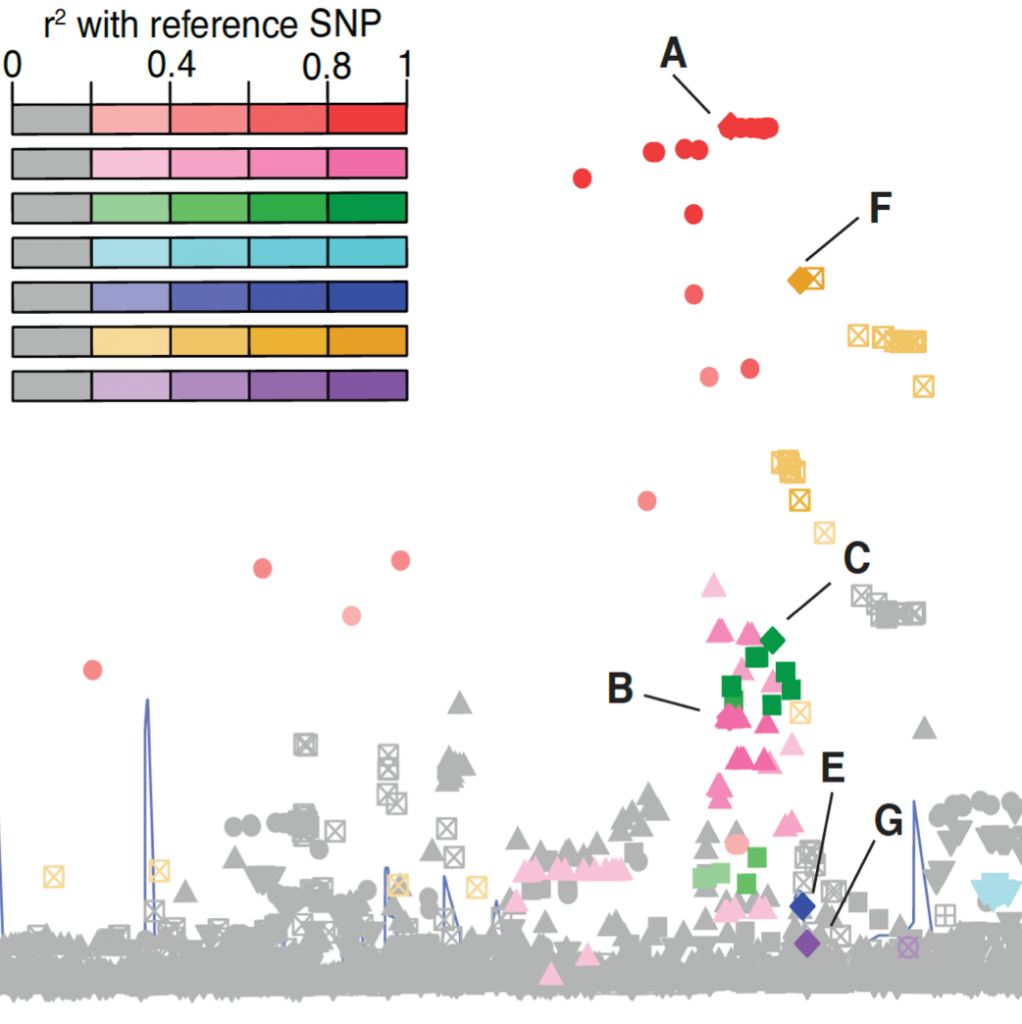
To identify genetic variants that confer disease susceptibility or influence variability in related traits, we conduct genome-wide association studies (GWAS) in human population studies. In recent years, we and our collaborators have used this approach to identify hundreds of new genetic loci associated with type 2 diabetes, obesity, cholesterol levels, metabolic traits, and cardiovascular risk factors. Large-scale meta-analyses and other multidisciplinary collaborations are key to many of these discoveries. We also study how genetic variants interact with environmental factors to influence the underlying biology of complex diseases and traits.
Characterize Disease- and Trait-Associated Variants
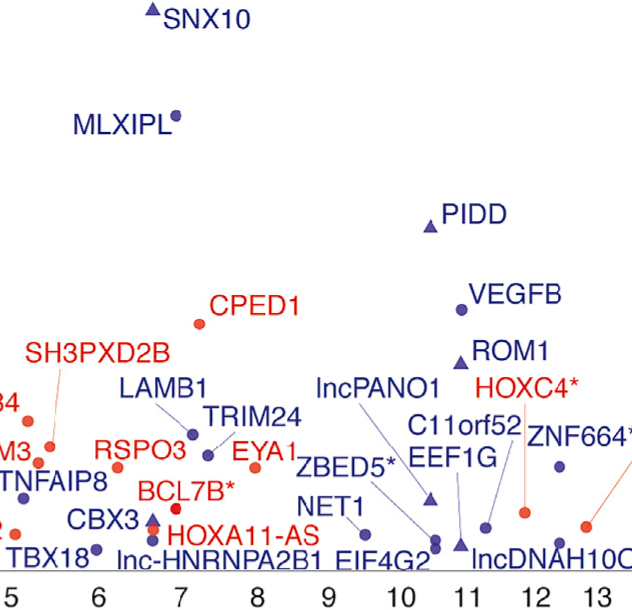

Loci identified by GWAS usually contain many variants in strong linkage disequilibrium with each other, and the genes responsible for most GWAS loci are unclear. We use computational approaches to identify candidate genes and candidate variants, especially at loci with putative regulatory mechanisms. Genome-wide data sets containing information about gene expression, chromatin structure, transcription factor binding, and epigenetic marks in human cells are important resources for identifying genes and regulatory regions. By generating and analyzing transcriptomic and epigenomic data sets, we identify regulatory elements and genes that vary due to genetic or environmental perturbation. We use these data to identify trait-associated variants likely to regulate gene regulation.
Investigate Molecular and Biological Mechanisms
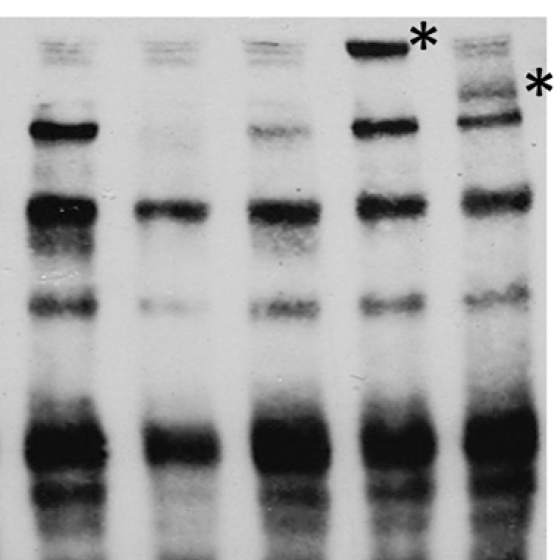
For most loci identified by association studies, the underlying molecular and biological mechanisms remain unknown. Determining the biological basis for the associations of genetic loci with complex diseases and traits will improve our general scientific understanding of pathways contributing to disease etiology. We employ molecular and cellular biology techniques such as CRISPR/Cas-based genome editing to identify the functional variant(s) and gene(s) at these loci, explain the molecular mechanisms linking functional variant(s) to gene(s), and characterize the biological mechanisms linking gene(s) to metabolic traits.